
zeae) es una enfermedad que se caracteriza porque el patógeno remplaza las inflorescencias por soros llenos de teliosporas. Key words: Zea mays native maize resistance incidence head smut artificial inoculationĮl carbón de la espiga del maíz ( Sporisorium reilianum f. Considering the geographical origin, the native maize collections from the Estados de Mexico and Tlaxcala, had a lower incidence of the disease compared to the rest of the populations, which indicates the presence of genes for resistance to the disease. 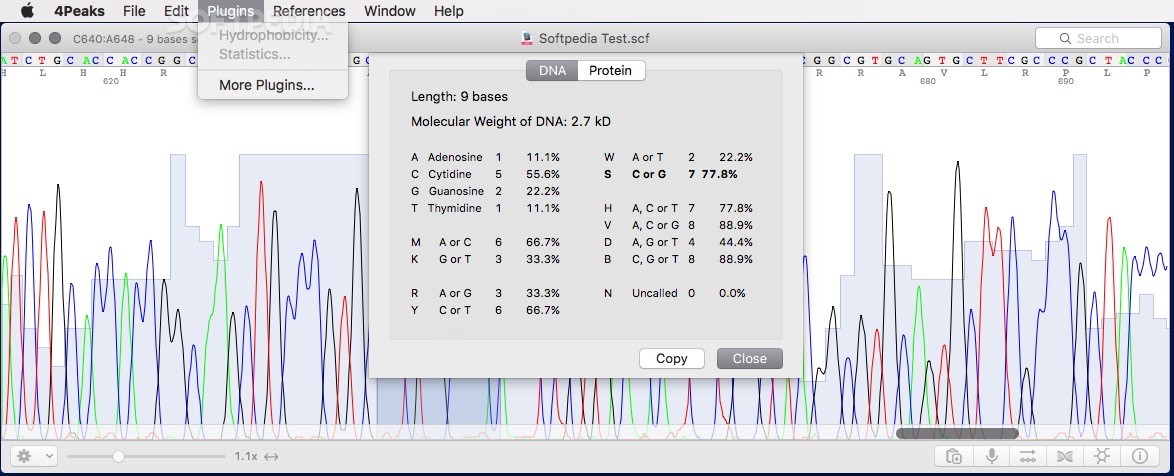
The maximum incidence of the disease in the maize populations was 28.8% and 22.2% in the first and second evaluation, respectively, while the control (Az 41801) presented an incidence of 70.7% and 42.3%. The incidence of the disease was recorded by direct observation of signs and symptoms in the inflorescences. The populations were evaluated in Mixquiahuala, Hidalgo, in the 20 plantings. The hybrid Az 41801 was used as a control. The seed was inoculated with teliospores of the pathogen, using grenetine as adherent. Maize populations were collected in the states of Guerrero (13), Oaxaca (13), Puebla (six), Tlaxcala (12) and Estado de México (11). The objective of this study was to investigate the response of 55 native maize populations to S. zeae) is a disease characterized by the pathogen replacing inflorescences with sori full of teliospores. Patient characteristics according to HER2 mutation status.Head smut of maize ( Sporisorium reilianum f. “4Peaks” was used to view and edit the sequence trace files ( ). The left half of Fig 3 shows the amplification curves of Eprobe-PCR, and the right half shows the electrogram of Sanger sequencing. The data were then transferred to Microsoft Excel (Microsoft, Redmond, WA, USA) and Cp values were evaluated.Įvaluation of the nine mutant tumors detected by Eprobe-PCR.Ĭomparison of three genotyping methods in all clinical samples.Ĭomparison of Eprobe-PCR and Sanger methods. (c) Cp (crossing point) values of two experiments (a) and (b) were calculated by the second derivative maximum method in the LightCycler480. The light blue line shows no amplification. The total copy number for each was adjusted to 10,000 copies per reaction. The blue line indicates MT only plasmid DNA at 10,000 copies per reaction, red: 1,000, green: 100, purple: 10, light blue: 1, orange: WT plasmid DNA, black: NTC (diluted water). (b) Sensitivity of 12-bp duplicated insertion detection in heterogenetic conditions. The blue line indicates MT only plasmid DNA at 10,000 copies per reaction, red: 1,000, green: 100, purple: 10, light blue: 1, orange: WT plasmid DNA, black: NTC. 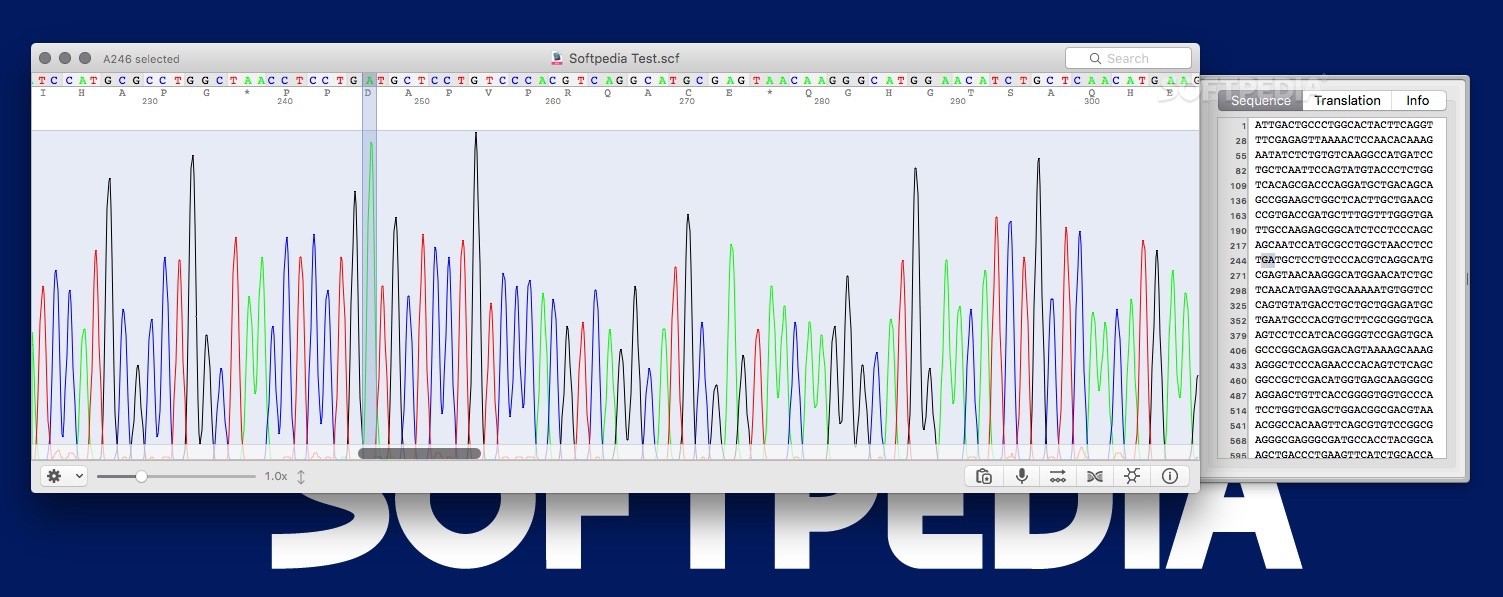
(a) Evaluation of mutated genome amplification. MT: HER2 12-bp duplicated insertion mutation type, WT: HER2 wild type, NTC: No template control (diluted water). Sensitivity of Eprobe-PCR for detecting HER2 12-bp duplicated insertion. The 3’ end-filled circle Eprobe shows the blocker that prevents primer extension during PCR. The forward primer for detection of the mutant-type allele contains the full sequence of HER2 across the region known to be a frequent insertion site. The orange box is the duplicated insertion. Schematic diagram of primers for the detection of the HER2 12-bp duplicated insertion by Eprobe-PCR. Primer sets and Eprobe design for HER2 mutation detection.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
